Introduction
The bgms package implements Bayesian methods for analyzing graphical models. It supports three variable types:
- ordinal (including binary) — Markov random field (MRF) models,
- Blume–Capel — ordinal MRF with a reference category,
- continuous — Gaussian graphical models (GGM).
The package estimates main effects and pairwise interactions, with optional Bayesian edge selection via spike-and-slab priors. It provides two main entry points:
-
bgm()for one-sample designs (single network), -
bgmCompare()for independent-sample designs (group comparisons).
This vignette walks through the basic workflow for ordinal data. For
continuous data, set variable_type = "continuous" in
bgm() to fit a Gaussian graphical model.
Wenchuan dataset
The dataset Wenchuan contains responses from survivors
of the 2008 Wenchuan earthquake on posttraumatic stress items. Here, we
analyze a subset of the first five items as a demonstration.
Fitting a model
The main entry point is bgm() for single-group models
and bgmCompare() for multiple-group comparisons.
fit = bgm(data, seed = 1234)Posterior summaries
summary(fit)
#> Posterior summaries from Bayesian estimation:
#>
#> Category thresholds:
#> mean mcse sd n_eff Rhat
#> intrusion (1) 0.486 0.005 0.233 2203.156 1.007
#> intrusion (2) -1.884 0.012 0.340 784.588 1.021
#> intrusion (3) -4.813 0.021 0.560 714.327 1.020
#> intrusion (4) -9.460 0.031 0.896 813.120 1.019
#> dreams (1) -0.601 0.003 0.190 3158.280 1.001
#> dreams (2) -3.799 0.007 0.343 2243.581 1.003
#> ... (use `summary(fit)$main` to see full output)
#>
#> Pairwise interactions:
#> mean mcse sd n_eff n_eff_mixt Rhat
#> intrusion-dreams 0.316 0.001 0.032 3935.279 1.000
#> intrusion-flash 0.168 0.000 0.031 3845.959 1.000
#> intrusion-upset 0.090 0.003 0.041 231.981 150.046 1.091
#> intrusion-physior 0.103 0.001 0.031 847.734 774.814 1.007
#> dreams-flash 0.250 0.000 0.030 5082.832 1.001
#> dreams-upset 0.116 0.001 0.029 1088.410 717.567 1.011
#> ... (use `summary(fit)$pairwise` to see full output)
#> Note: NA values are suppressed in the print table. They occur here when an
#> indicator was zero across all iterations, so mcse/n_eff/n_eff_mixt/Rhat are undefined;
#> `summary(fit)$pairwise` still contains the NA values.
#>
#> Inclusion probabilities:
#> mean mcse sd n0->0 n0->1 n1->0 n1->1 n_eff_mixt
#> intrusion-dreams 1.000 0.000 0 0 0 3999
#> intrusion-flash 1.000 0.000 0 0 0 3999
#> intrusion-upset 0.889 0.032 0.314 425 18 18 3538 93.525
#> intrusion-physior 0.992 0.006 0.090 29 4 4 3962 260.352
#> dreams-flash 1.000 0.000 0 0 0 3999
#> dreams-upset 0.996 0.004 0.067 16 2 2 3979
#> Rhat
#> intrusion-dreams
#> intrusion-flash
#> intrusion-upset 1.205
#> intrusion-physior 1.052
#> dreams-flash
#> dreams-upset 1.086
#> ... (use `summary(fit)$indicator` to see full output)
#> Note: NA values are suppressed in the print table. They occur when an indicator
#> was constant or had fewer than 5 transitions, so n_eff_mixt is unreliable;
#> `summary(fit)$indicator` still contains all computed values.
#>
#> Use `summary(fit)$<component>` to access full results.
#> Use `extract_log_odds(fit)` for log odds ratios.
#> See the `easybgm` package for other summary and plotting tools.You can also access posterior means or inclusion probabilities directly:
coef(fit)
#> $main
#> cat (1) cat (2) cat (3) cat (4)
#> intrusion 0.4863539 -1.883558 -4.813051 -9.460128
#> dreams -0.6005757 -3.798926 -7.131576 -11.567385
#> flash -0.1041094 -2.572478 -5.376715 -9.684421
#> upset 0.4228417 -1.289224 -3.341700 -6.985633
#> physior -0.6194550 -3.181700 -6.234517 -10.589579
#>
#> $pairwise
#> intrusion dreams flash upset physior
#> intrusion 0.00000000 0.3157417314 0.168320925 0.090174534 0.1034944713
#> dreams 0.31574173 0.0000000000 0.249999922 0.116103200 0.0003192306
#> flash 0.16832093 0.2499999222 0.000000000 0.005532944 0.1525726218
#> upset 0.09017453 0.1161031996 0.005532944 0.000000000 0.3548717664
#> physior 0.10349447 0.0003192306 0.152572622 0.354871766 0.0000000000
#>
#> $indicator
#> intrusion dreams flash upset physior
#> intrusion 0.00000 1.00000 1.0000 0.88925 0.99175
#> dreams 1.00000 0.00000 1.0000 0.99550 0.00775
#> flash 1.00000 1.00000 0.0000 0.10450 1.00000
#> upset 0.88925 0.99550 0.1045 0.00000 1.00000
#> physior 0.99175 0.00775 1.0000 1.00000 0.00000Network plot
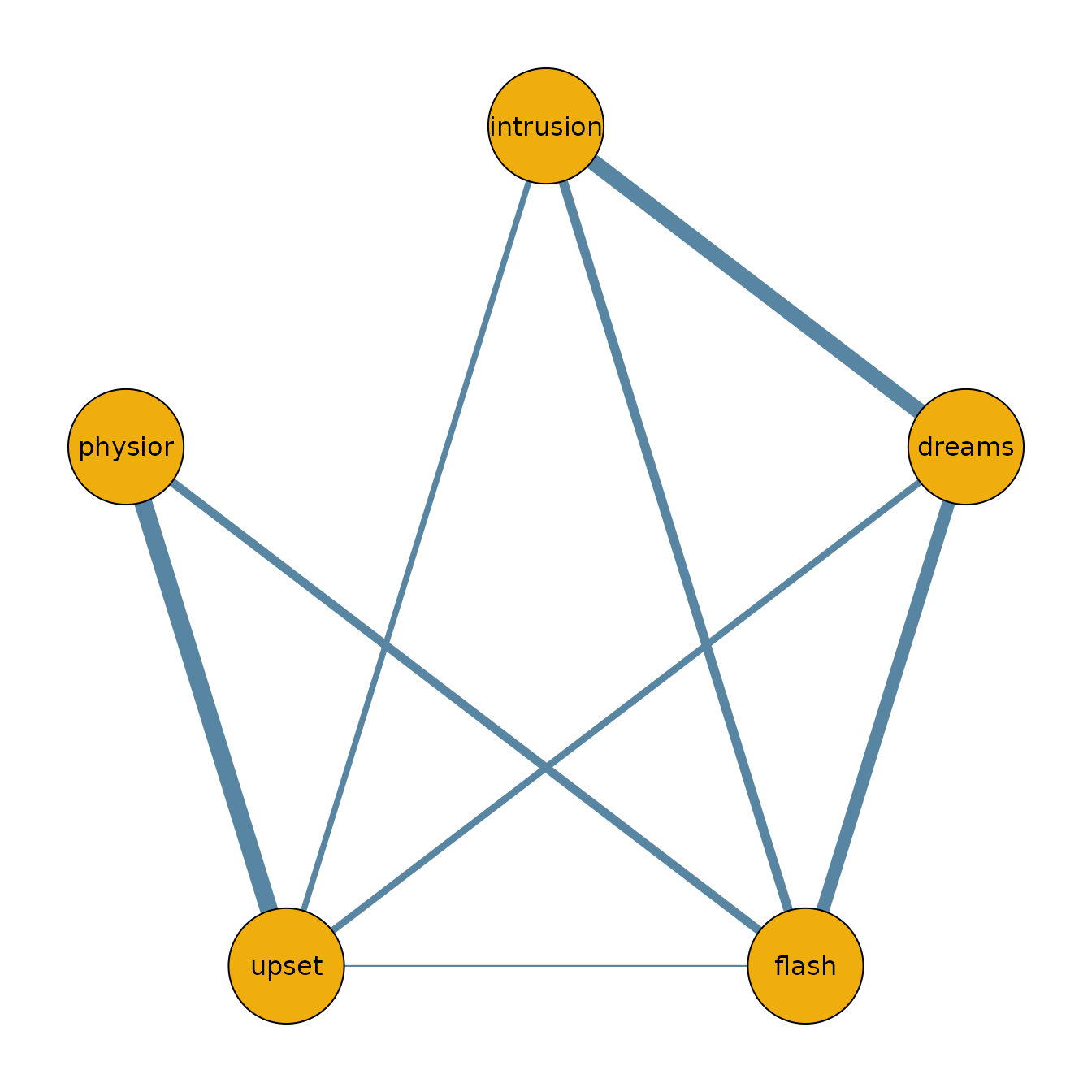
To visualize the network structure, we threshold the posterior inclusion probabilities at 0.5 and plot the resulting adjacency matrix.
library(qgraph)
median_probability_network = coef(fit)$pairwise
median_probability_network[coef(fit)$indicator < 0.5] = 0.0
qgraph(median_probability_network,
theme = "TeamFortress",
maximum = 1,
fade = FALSE,
color = c("#f0ae0e"), vsize = 10, repulsion = .9,
label.cex = 1, label.scale = "FALSE",
labels = colnames(data)
)
Continuous data (GGM)
For continuous variables, bgm() fits a Gaussian
graphical model when variable_type = "continuous". The
workflow is the same:
The pairwise effects are partial correlations (off-diagonal entries
of the standardized precision matrix). Missing values can be imputed
during sampling with na_action = "impute".
Next steps
- For comparing groups, see
?bgmCompareor the Model Comparison vignette. - For diagnostics and convergence checks, see the Diagnostics vignette.